Molecular Parasitology (Prof. A. Hehl)
Parasites rule!
Toxoplasma gondii, a successful parasite
Toxoplasma gondii is a globally prevalent zoonotic parasite from the phylum Apicomplexa. On the one hand, T. gondii is an important cause for abortions in sheep, on the other hand approximately one third of the human population is infected with T. gondii resulting in considerable morbidity. T. gondii is capable of infecting any warm-blooded animal as intermediate host, where after a period of acute infection mediated by tachyzoites the parasite can persist as a cyst containing slow-replicating bradyzoites. Chronic toxoplasmosis remains untreatable and can revert to an acute infection upon immunosuppression causing a stage-conversion of bradyzoites to tachyzoites. This can yield among others to encephalitis and chorioretinitis.
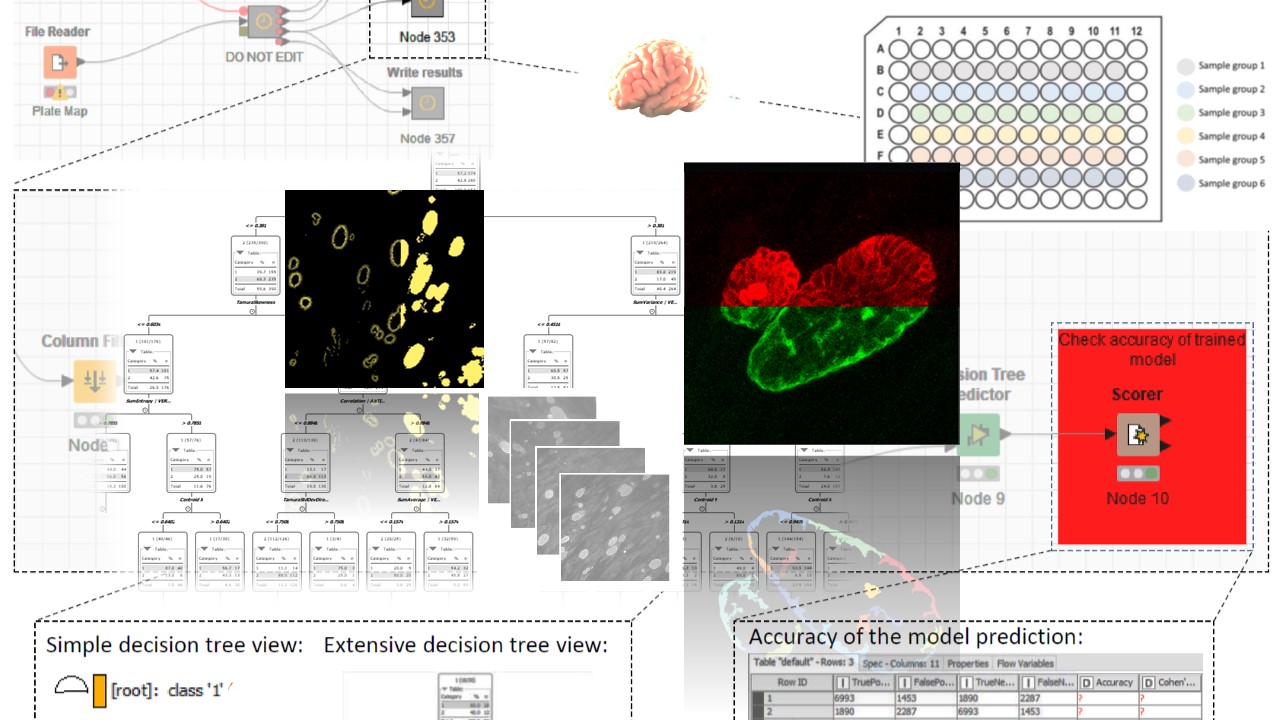
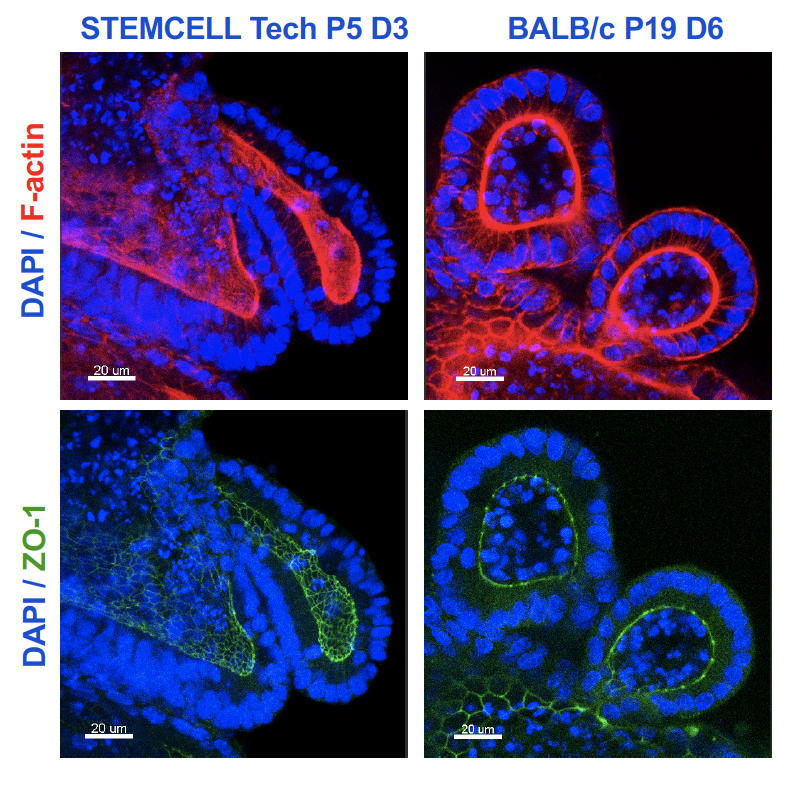
In a collaborative effort, our lab works on the elucidation of structure and biogenesis of the tissue cyst wall of T. gondii but also of related coccidia such as Besnoitia besnoiti, a parasite of cattle. We exploit the well-stocked molecular genetic toolbox to analyze loss-of-function mutant phenotypes in in vitro cultivated tissue cysts. A reproducible high-throughput, high-content fluorescence microscopy and image analysis pipeline has been established for the characterisation of tissue cyst morphology and cyst wall aberrations in loss-of-function mutants. Our interests are primarily in factors and processes related to export to and across the cyst wall as well as communication across this biological barrier
The main focus of our lab is the cell biology of developmental stages in the felid definitive host. Using deep RNA sequencing of parasites harvested from experimental infections, we established the baseline for developmental regulation of gene expression during the asexual expansion (merozoite stages) and the development of sexual stages (gametocytes) and zygotes in cat intestinal epithelial cells. With the elucidation of developmental regulation of mRNA expression during the external sporulation of oocysts we now have a documentation of the complete
T. gondii life cycle.
Giardia lamblia, an almost perfect parasite
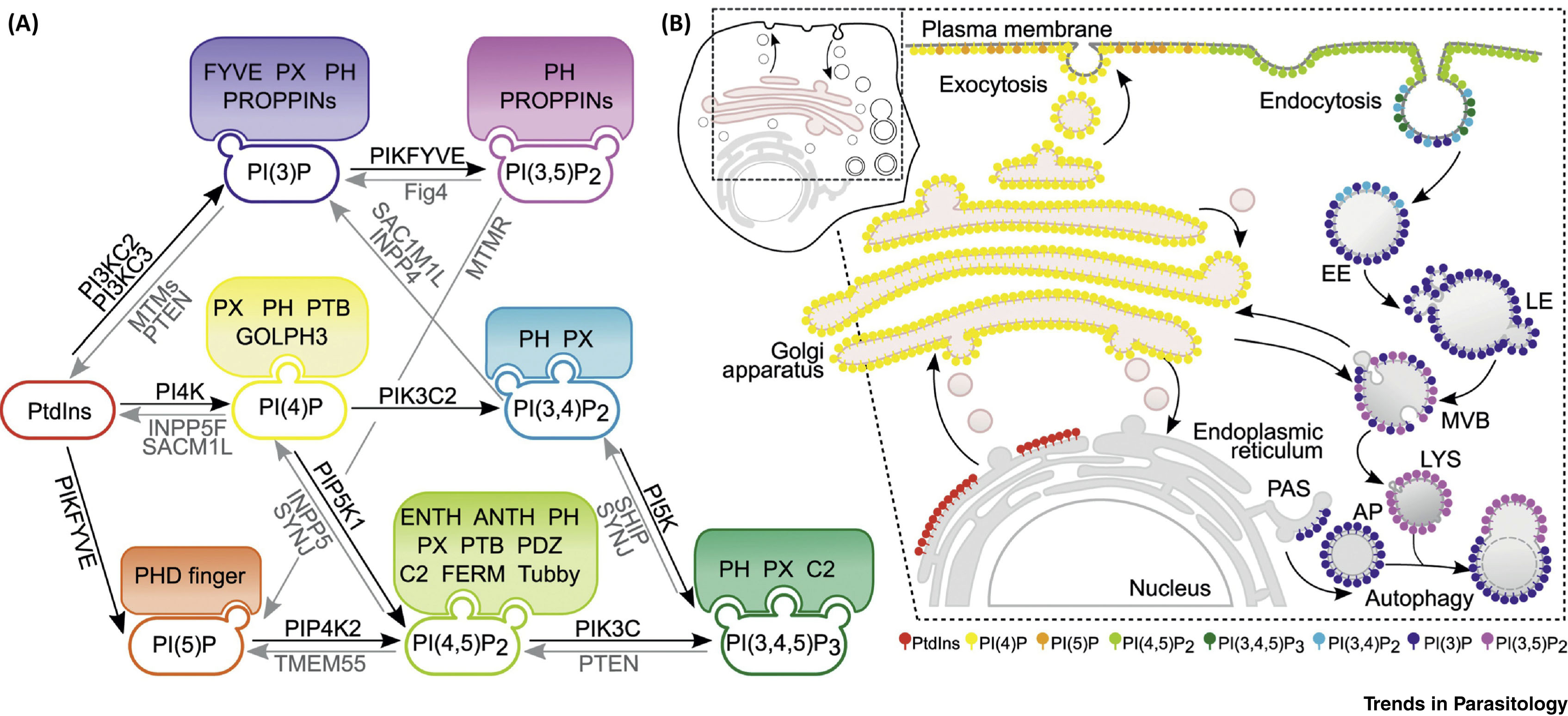
Over the past years, our laboratory has investigated many aspects of the subcellular organization of the Excavata member Giardia lamblia (syn. duodenalis, intestinalis) in considerable detail. There are several reasons for this endeavour, which transcend this parasite's medical importance, which was primarily motivated by the reduced subcellular complexity and debated evolutionary status of this parasite. One may say that simplification/loss of complexity or “reverse” evolution has emerged as a paradigm for the evolution of Giardia's subcellular architecture. However, a complete appreciation of the evolutionary and ecological significance of this phenomenon is far from complete. Our experimental research started out with elucidating membrane trafficking pathways for protein export in G. lamblia in the absence of a Golgi structure or function(s) and expanded to include endo- and exo-cytosis, organellar protein import, morphology and function. We also addressed questions concerning organelle replication and inheritance while strongly driving technical advances, in particular new methods and approaches to genetic manipulations in G. lamblia. By providing a basic framework for protein trafficking and organelle genesis, we gained a better understanding of the G. lamblia subcellular organization at the morphological and molecular level. This work is complemented by efforts aimed at elucidating this parasitic species' evolutionary status and could provide us with the basis for novel strategies to interfere with parasite transmission and/or pathogenesis.
As of 2019, this research on G. lamblia has moved to the laboratory of Dr. Carmen Faso at the Institute of Cell Biology of the University of Bern
(https://www.izb.unibe.ch/research/dr_carmen_faso/index_eng.html#pane823287).
The Laboratory for Molecular Parasitology is at the Institute of Parasitology which belongs to the Vetsuisse Faculty and the Faculty of Medicine of the University of Zürich and is affiliated with the Life Sciences Zürich Graduate School.
Metabolism and Subcellullar Localization of Phosphoinositides (PIPs) in Mammalian Cells.